Introduction¶
When I started my PhD at the Erasmus MC in 2016 on radiomics, the idea was to start with a straight-forward radiomics study to get familiar with the field. Both the field of radiomics and my group at the Erasmus MC had existed for quite some time. Therefore, I naively expected that all previously created methods and software were nicely organized and would be straight forward to apply and compare. However, I found that of all the radiomics studies, only a few actually released their software open-source. Moreover, the software that was available was very diverse in terms of programming language and often difficult to understand. Each developer used its own ontology, workflow and implementation.
Hence I wondered which method would work best for my specific application. It seemed that many of my fellow PhD students started out with the same question. I did not like the foresight to start out my PhD by tediously working on finding a decent baseline and trying out all kinds of methods. Moreover, it felt inefficient that every time a new study is started, someone like me had to go through the whole process over and over again.
I therefore decided to create one radiomics platform in which multiple approaches could easily be integrated. I tried to make it as flexible as possible and modular, such that researcher can easily integrate their own components into the workflow.
Including all these methods brought along many choices in terms of both the methods to use and their parameters. Similar to the above, I also did not like the foresight of manual tuning, which both feels inefficient and a non-optimal approach. Hence, I decided to create an automatic optimization routine for the platform which includes all the choices in the workflow that have to be made.
The result is WORC, which has made my research a lot easier and more fun, as I can very easily create a decent radiomics baseline for any study with little effort.
Radiomics Terminology¶
Radiomics concerns the use of quantitative medical image features to predict clinically relevant outcomes. In WORC, the computational radiomics workflow is divided in five steps:
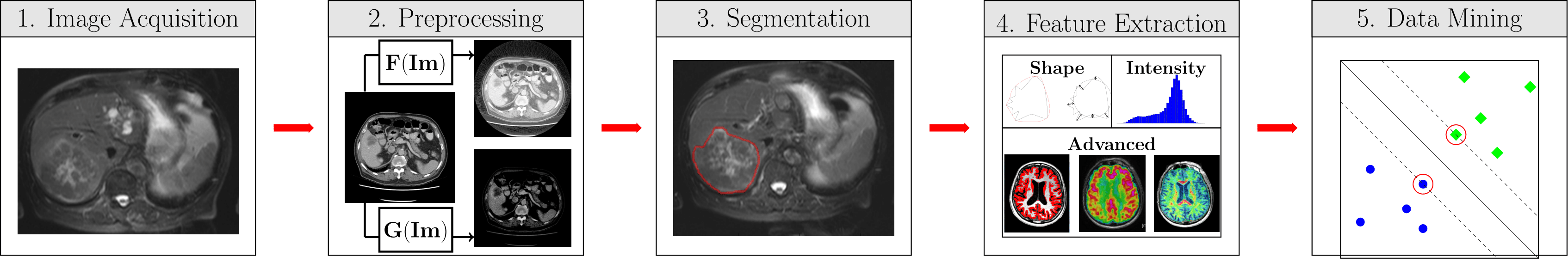
Image Acquisition, which we assume has already been done.
Preprocessing
Segmentation
Feature Extraction
Data Mining
Preprocessing is defined as anything that is done to the image before the feature extraction. This may include normalization, cropping, masking, and registration. Filtering may be included as well, although several features may already implicitly use a filter.
Data mining is defined as anything that is used to go from features to the actual label or outcome. This may include feature selection, feature imputation, feature scaling, dimensionality reduction, and oversampling. Note that we always define the last step in data mining as a machine learning approach. Although several radiomics studies do not use machine learning but directly correlate features with outcome, we feel that in clinical practice a prediction model is most of the time required.
Due to the shear amount of tools available in the data mining node, this node in itself is another fastr (micro)network.
Modularity and standardization: fastr¶
The first step of building WORC was to standardize radiomics in components to create a modular workflow. The available radiomics methods were all in different software languages, hence my framework had to support this as well. Luckily, some of my colleagues had the same issues and decided to create a new workflow engine: fastr. I highly recommend to read at least the introductory page of the manual in order to get familiar with the basic concept of fastr .
To illustrate the principles of fastr, one of the simpler workflows created by WORC using fastr is shown in the figure below.
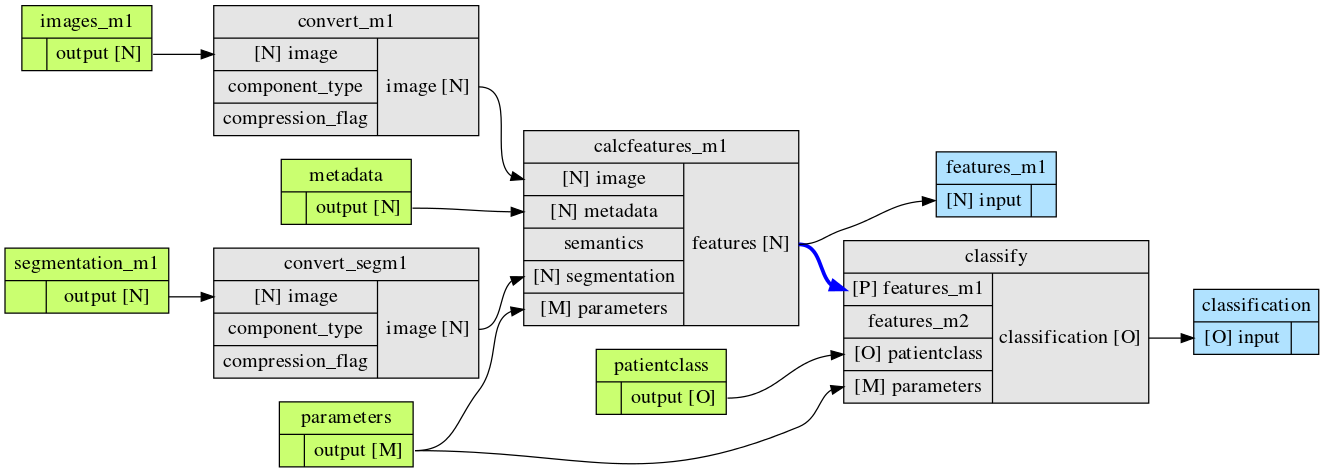
The radiomics workflow is split in nodes. We define four types of nodes: * Sources, which serve as the input for the workflow. * Nodes, which is where the actual algorithms run. * Sinks, which serve as the output for the workflow. * Constants
In each node, the inputs, outputs and thereby the task is fixed. However, the actual algorithm that is used within a node is node. For example, in a feature extraction node, the inputs are an image, possibly a segmentation and parameters, from which features are extracted, which are thereby the output. Which methods is actually used to extract the features and the features extracted may vary.
Using fastr in WORC gives us several benefits: * You can use any tool that can be run on the command line: thus languages or interpreters like MATLAB, Python, R, but also executables. * As the workflow is standardized, the outputs are as well. Hence workflows can be easily compared. * fastr automatic keeps a history of how an output was created, or the provenance, and extensive logging. This facilitates high reproducability. * Wrapping a tool in fastr is easy: you only need to determine the inputs, outputs, interpreter and script. * fastr supports various execution plugins, thus execution of a pipeline on a cluster or a simple laptop can be adapted to the resources at hand. * Data can be imported and exported to various sources, also online!
Optimization¶
When starting a new radiomics study, e.g. trying to find a imaging biomarker on a new dataset, you would normally have to design a workflow yourself from all methods available in WORC. Not only would you have to determine which methods you are going to use, but also which parameters for all methods. Many studies have shown that these are often not independent: the performance of many methods highly depends on the parameters used. Thus, we need to simultanously look for the best combination of these.
As these parameters are set before the actual learning step in the machine learning part, we refer to these as hyperparameters. As the performance of the machine learning algorithm may depend on the previous steps (e.g. preprocessing, feature extraction, and all other steps in the data mining node), we decided to include all parameters from all nodes as hyperparameters.
Optimization is done through a variant of combined algorithm selection and hyperparameter (CASH) optimization problem. Currently, WORC solves the problem as following:
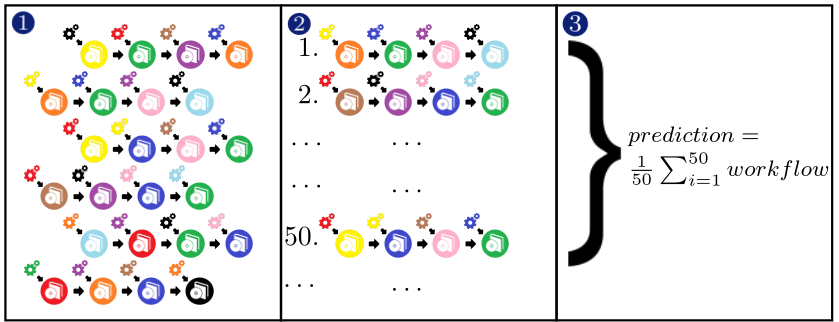
Generate a large number of different pipelines.
Execute them and rank them.
Create and ensemble of the 50 best performing pipelines.